[Prävalenz von Helicobacter pylori-Infektionen bei Patienten mit Gastritis und Dyspepsie in Jos, Bundesstaat Plateau, Nigeria]
David Veronica Ekpiwre 1Hashimu Zakari 1
Anayochukwu Chibuike Ngene 2
Hosea J. Zumbes 1
Amos Obaje Ogaji 1
Oluwatoyin Debby Coulthard 1
John Danjuma Mawak 1
1 Department of Microbiology, Faculty of Natural Sciences, University of Jos, Jos, Plateau State, Nigeria
2 Department of Microbiology, College of Natural Sciences, Michael Okpara University of Agriculture, Umudike, Nigeria
Zusammenfassung
Hintergrund: Eine Infektion mit Helicobacter (H.) pylori ist weltweit nach wie vor eine der Hauptursachen für Gastritis und Dyspepsie, wobei die Prävalenz in Entwicklungsregionen unverhältnismäßig hoch ist, darunter auch in Teilen Afrikas mit Infektionsraten oft über 70%. In vielen dieser Regionen behindern begrenzte regionsspezifische Daten zur Prävalenz und Antibiotikaresistenz die wirksame Diagnose, Behandlung und Bekämpfungsstrategien.
Zielsetzung: Untersucht werden sollten die Prävalenz, die molekularen Eigenschaften und die Antibiotikaempfindlichkeit von H. pylori bei Patienten mit Gastritis und Dyspepsie in Jos im Bundesstaat Plateau, Nigeria.
Methoden: Es wurden 136 symptomatische Patienten rekrutiert, 36 mit Dyspepsie, die sich einer Endoskopie unterzogen, und 100 mit Gastritis, die Stuhlproben zur Verfügung stellten. Die Proben (Magenbiopsien und Stuhl) wurden zunächst mit einem Urease-Schnelltest (RUT) und einem Stuhl-Antigentest (SAT) untersucht. Positive Proben wurden zur Isolierung kultiviert, anschließend phänotypisch identifiziert und mittels Polymerasekettenreaktion (PCR) mit genus- und artspezifischen Primern bestätigt. Die Antibiotikaempfindlichkeit wurde mittels Disk-Diffusionsmethode bewertet.
Ergebnisse: Die Prävalenz von H. pylori betrug 66,7% bei Patienten mit Dyspepsie und 48% bei Patienten mit Gastritis. Die Infektionsraten standen in beiden Gruppen in signifikantem Zusammenhang mit dem Alter (p<0,05), jedoch nicht mit dem Geschlecht. Die Kultur ergab 10 mutmaßliche Helicobacter-Isolate; mittels PCR wurden 6 als Helicobacter-Spezies und 3 als H. pylori bestätigt. Die Empfindlichkeitstests zeigten eine hohe Resistenz gegenüber Ciprofloxacin, Clarithromycin, Azithromycin und Amoxicillin. Alle Isolate waren empfindlich gegenüber Streptomycin mit unterschiedlichen Reaktionen auf Ofloxacin, Augmentin (Amoxicillin-Clavulanat) und Septrin (Trimethoprim-Sulfamethoxazol), was auf das Auftreten multiresistenter Stämme hinweist.
Schlussfolgerung: Die Ergebnisse zeigen eine erhebliche H. pylori-Belastung bei symptomatischen Patienten in Jos, verbunden mit einer alarmierenden Resistenz gegen wichtige Antibiotika zur Eradikation und unterstreichen die dringende Notwendigkeit einer routinemäßigen molekularen Diagnostik und kontinuierlichen lokalen Überwachung der Antibiotikaresistenz, um maßgeschneiderte Behandlungsschemata zu entwickeln und die klinischen Ergebnisse zu verbessern.
Schlüsselwörter
Helicobacter pylori, Gastritis, Dyspepsie, Prävalenz, Antibiotikaesistenz, Nigeria
Introduction
Helicobacter (H.) pylori infection continues to pose a significant public health threat owing to its close links with chronic gastritis, dyspepsia, peptic ulcers, and gastric cancers [1], [2]. The pathogen spreads mainly via fecal-oral and oral-oral transmission through contaminated food or water, or direct person-to-person contact, which promotes rapid dissemination in areas with poor hygiene and sanitation [3], [4], [5]. Its successful colonization of the gastric environment relies on key adaptive features, such as flagellar motility, urease enzyme activity to counteract stomach acid, and adhesins that enable firm attachment to epithelial cells, thereby sustaining long-term infection and ongoing mucosal inflammation [6], [7].
Worldwide, H. pylori affects roughly half the global population, with markedly higher rates (often reaching 80–90%) in low- and middle-income countries, including Nigeria [8], [9]. The bacterium has been designated a class I carcinogen, playing a major role in cancers attributable to infectious agents [10], [11]. Considerable genetic and phenotypic diversity among strains adds complexity to disease manifestation, host responses, and therapeutic success [12].
A pressing issue in treating H. pylori is the escalating antimicrobial resistance [13]. Resistance to standard agents in eradication therapies (such as clarithromycin, metronidazole, amoxicillin, and fluoroquinolones) is rising globally, resulting in reduced cure rates and higher recurrence [14]. In resource-limited settings like Nigeria, treatment is often prescribed empirically without localized data on infection rates or resistance profiles, potentially driving the emergence of multidrug-resistant strains and contributing to treatment failures [15], [16].
To address these gaps, localized studies on prevalence, molecular traits, and antibiotic susceptibility of H. pylori in symptomatic individuals are essential [17], [18]. This study investigates the infection prevalence among patients with gastritis and dyspepsia, while examining the phenotypic and genotypic properties of isolated strains, with a focus on resistance patterns and multidrug resistance. The results aim to guide evidence-based treatment choices, strengthen antimicrobial stewardship efforts, and optimize clinical management of H. pylori infection in the target population.
Materials and methods
Study population
A total of 36 dyspeptic patients experiencing upper abdominal pain or discomfort, bloating or fullness after meals, nausea and vomiting, heartburn or acid reflux attending the endoscopy unit of Jos University Teaching Hospital (JUTH) and 100 gastritis patients experiencing severe stomach pain and discomfort, and some with bleeding from the stomach attending Plateau State Specialist Hospital in Jos North LGA of Plateau State of different ages were included.
Inclusion and exclusion criteria
Patients with either Dyspepsia or gastritis who consented were included. While those with without dyspepsia or gastritis and those with who did not consent were excluded.
Ethical consideration
Ethical clearance was obtained from the Ethics Committee of the Jos University Teaching Hospital, Jos (REF: JUTH/DCS/IREC/127/XXXI/424). Informed consent was sought and obtained from all participants.
Isolation of H. pylori from biopsy specimens
Gastric biopsy samples were obtained from 36 dyspeptic patients undergoing endoscopy at the Endoscopy Unit of Jos University Teaching Hospital (JUTH). The specimens were immediately transferred into rapid urease broth for initial screening [19], [20]. Only biopsies yielding a positive rapid urease test were selected for bacterial isolation. These were cultured on brain heart infusion (BHI) agar plates supplemented with 7% sheep blood. The inoculated plates were incubated at 37°C in a microaerobic atmosphere to provide optimal growth conditions. Plates were examined daily for colonial growth starting from day 3 up to day 12.
Isolation of H. pylori from stool
Fresh fecal samples were collected in a sterile wide-mouthed air-tight containers from 100 gastritis patients. The samples were then inoculated into BHI broth and transported to the laboratory under cold chain (ice packs). The stool samples were first processed using H. pylori stool antigen (HpSA) kit. The H. pylori positive stool samples were emulsified in BHI broth and then incubated at 37°C for 24 hrs under anaerobic environment specifically in carbon dioxide extinction candle jar. Positive broth samples were then cultured on BHI agar supplemented with antibiotics which included 10 mg vancomycin and 5 mg Amphotericin B with 7% sheep blood. The plates were then incubated at 37°C under anaerobic environment. The agar plates were then checked for growth from day 3 through day 12 [21].
Preliminary identification of H. pylori
The inoculated plates were incubated in 100% humidity at 37°C for up 3–12 days in microaerophilic condition. Spot test using oxidase was then conducted on the colonies to check for the presence of suspected H. pylori before being sub-cultured on nutrient agar. An isolate was identified as H. pylori on the basis of positive catalase, oxidase and urease reaction, typical colonial morphology of small, round and greyish colonies and the presence of characteristic curved Gram-negative short rods on Gram Stain [22]. Polymerase chain reaction (PCR) was done to further confirmed the isolates.
Gram staining
On a clean grease free slide, a small drop of normal saline was placed. Using a sterile wire loop a discreet colony from the cultured plate was picked and emulsified on the slide to make a thin smear and was allowed to air-dry. The smear was heat fixed using dry heat. The heat fixed smear was stained with crystal violet (primary stain) and allowed to stay for 60 seconds. The smear was rinsed with water and floated with Lugol's iodine solution (mordant) then allowed for 30 seconds. The iodine was rinsed off with water. Using acetone (decolorizing agent), the smear was flooded for 5 seconds and rinsed immediately under running water. This generates the differential aspect of Gram staining. Finally, smear was counter-stained with safranin (secondary stain) for 60 seconds, rinsed off under running water and the slide was allowed to air-dry. The slide was viewed under the microscope using ×100 oil immersion objectives. H. pylori were seen as Gram-negative curve rods [23].
Oxidase test
A drop of 1% oxidase reagent was applied to a strip of filter paper. A portion of a colony from the culture plate was collected using a sterile wire loop and rubbed onto the reagent-impregnated area. Development of a deep purple coloration within 20 seconds signified a positive result [23].
Catalase test
On a clean, grease-free glass slide, a colony from the culture plate was transferred using a sterile inoculating loop and smeared across the surface. A drop of 3% hydrogen peroxide was added to the smear and gently mixed. The immediate formation of gas bubbles indicated a positive catalase reaction [23].
Urease test
Using a flamed wire loop, a colony was picked from the culture plate and stabbed on urease agar slant inside test tube, it was covered and incubated at 37°C for 72 hrs. A color change was observed from orange to pink following incubation which indicated positive test [23].
Maintenance of H. pylori
After purification and identification of the clinical isolates of H. pylori, strains were preserved in a small screw capped tube in BHI broth at –4°C.
Antibiotic susceptibility testing
Antibiotic susceptibility testing was conducted using the disk diffusion method, in accordance with previously described protocols [24], [25] and the Clinical and Laboratory Standards Institute (CLSI) guidelines [24]. A single isolated colony was emulsified in peptone water and adjusted to a turbidity matching the 0.5 McFarland standard. The bacterial suspension was then evenly inoculated onto BHI agar plates. Antibiotic disks were applied to the agar surface, and the plates were incubated under microaerobic conditions at 37°C for 72 hrs. Inhibition zone diameters were measured and interpreted to classify isolates as susceptible, intermediate, or resistant. The antibiotics tested, along with their disk potencies, included ofloxacin (tarivid, 10 µg), trimethoprim-sulfamethoxazole (septrin, 30 µg), chloramphenicol (30 µg), nalidixic acid (30 µg), ciprofloxacin (10 µg), amoxicillin (10 µg), amoxicillin-clavulanate (augmentin, 30 µg), gentamicin (10 µg), pefloxacin (preflacine, 10 µg), and streptomycin (30 µg) (agents frequently employed in standard H. pylori eradication regimens).
Molecular identification
Presumptive H. pylori isolates were subjected to confirmatory identification via polymerase chain reaction (PCR) analysis. Genomic DNA was extracted from phenotypically identified H. pylori isolates using the QIAamp DNA Mini Kit (Qiagen), in accordance with the manufacturer’s instructions. Briefly, H. pylori colonies were collected using a sterile plastic loop and suspended in 200 µL of phosphate-buffered saline (PBS) in sterile 1.5 mL microcentrifuge tubes. The suspensions were vortex-mixed and centrifuged at 8,000 rpm for 1 minute. The supernatants were discarded, and the pellets were resuspended in a fresh 200 µL volume of PBS by vortexing. The resuspended cells were transferred into new sterile 1.5 mL microcentrifuge tubes containing 20 µL of proteinase K, followed by the addition of 200 µL of Buffer AL. The mixtures were thoroughly vortexed for 15 seconds and incubated at 56°C for 10 minutes to achieve cell lysis. The tubes were then briefly centrifuged at 8,000 rpm to remove condensation from the lids. The lysates were applied to QIAamp Mini spin columns placed in 2 mL collection tubes and centrifuged at 8,000 rpm for 1 minute. The columns were transferred to fresh collection tubes, washed with 500 µL of Buffer AW1, and centrifuged at 8,000 rpm for 1 minute, after which the flow-through was discarded. A second wash was performed using 500 µL of Buffer AW2, followed by centrifugation at 14,000 rpm for 3 minutes. To prevent buffer carryover, an additional dry spin was performed at 14,000 rpm for 1 minute. Finally, the spin columns were placed into sterile 1.5 mL microcentrifuge tubes, and 200 µL of Buffer AE (elution buffer) was added directly onto the membrane. After incubation at room temperature for 1 minute, the columns were centrifuged at 8,000 rpm for 1 minute to elute the DNA. The extracted DNA was used immediately for PCR amplification.
PCR amplification and gel electrophoresis
The following primers were used: HELF: 5’-AACGATGAAGCTTCTAGCTTGCTA-3’, HELR: 5’-GTGCTTATTCSTNAGATACCGTCAT-3’ [17], HPYF: 5’-GCGACCTGCTGGAACATTAC-3’ and HPYR: 5’-CGTTAGCTGCATTACTGGAGA-3’ [18]. Amplification of bacterial DNA was done using 25 µl total reaction volume containing 12.5 µl of 2X master mix, 1 µl of forward primer, 1 µl reverse primer and, 5.5 µl of nuclease-free water and 5 µl of DNA each for Helicobacter spp. specific (HEL) and H. pylori specific (HPY) detections. The thermocycling profile for HEL was as follows: Initial denaturation at 94°C for 5 min, followed by 35 cycles of denaturation at 94°C for 1 min, annealing at 56°C for 1 min and extension at 72°C for 1 min, held for final extension at 72°C for 10 min. The expected band size was 399 bp. For HPY, each reaction mixture was amplified for 35 cycles as follows: 30 sec at 94°C, 30 sec at 55°C and 30 sec at 70°C. A preincubation of 5 min at 94°C and final extension of 72°C for 30 sec were performed. The expected band size was 138 bp. The amplified products were visualized on 1.5% agarose gel stained with 8 µl ethidium bromide and was ran for 35 min at 100 volts.
Results
The overall prevalence of H. pylori in this dyspeptic cohort was 66.7% (24 out of 36 patients) (Table 1 [Tab. 1]). A significant association was found between age group and H. pylori occurrence (Χ²=7.679, p-value=0.022). The infection rate was highest in the 30–39 years age group (85%, 17 out of 20), followed by the 40–49 years group (66.7%, 2 out of 3). The lowest prevalence was observed in the 50–59 years age group (38.5%, 5 out of 13). In contrast, no statistically significant association was found between gender and H. pylori infection (Χ²=0.056, p-value=0.813).
Table 1: Occurrence of H. pylori in cases of dyspeptic patients in relation to age and gender (n=36)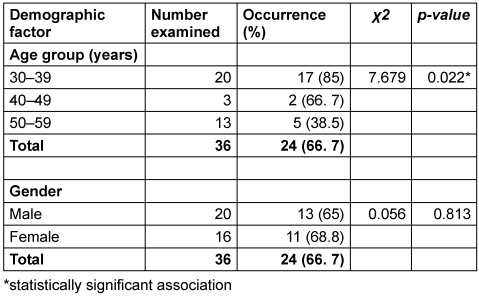
Table 2 [Tab. 2] presents the distribution of H. pylori prevalence in stool specimens from gastritis patients, stratified by age and gender (n=100). The overall infection rate was 48%. A statistically significant association was observed between age and H. pylori positivity (Χ²=14.336, p=0.006). The highest prevalence occurred in the 50–59 years age group (76.2%; 16/21), followed closely by the 20–29 years group (61.9%; 13/21). In contrast, the lowest rate was recorded among patients aged 60 years and older (27.3%; 6/22). Regarding gender, prevalence was higher among females (52.8%; 38/72) than males (35.7%; 10/28), although complete statistical details for this comparison were not specified in the table.
Table 2: Prevalence of H. pylori in the stool of gastric patients in relation to age and sex (n=100)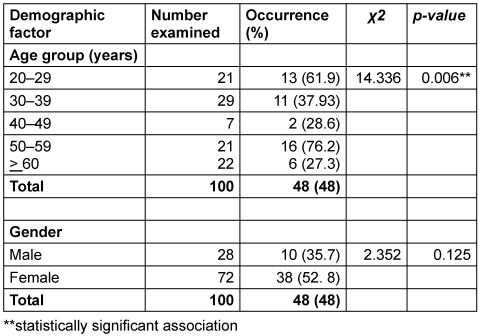
The occurrence of H. pylori and other isolates of the culture medium from dyspeptic and gastric patients is presented in Table 3 [Tab. 3]. A total of 75 isolates were obtained of six different genera were identified. Escherichia (E.) coli had the highest occurrence at 36%, followed by Pseudomonas species, which accounted for 26.7% of the isolates. H. pylori appeared in 13.3% of samples, while yeast cells were detected in 10.7% of the samples. Bacillus species showed a lower frequency of 8%, and Acinetobacter species had the least occurrence at 5.3%.
Table 3: Occurrence of H. pylori and other organisms in dyspeptic and gastric patients (n=75)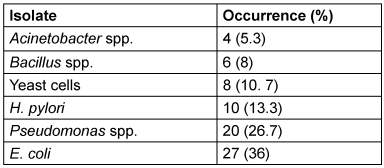
Molecular confirmation of presumptive Helicobacter species isolates
PCR analysis using Helicobacter genus-specific primers (HELF/HELR) produced the expected amplicon of approximately 399 bp in six of the ten presumptive isolates, confirming their identity as Helicobacter species. The molecular weight marker showed clear reference bands, while the negative control exhibited no amplification, indicating the absence of contamination or nonspecific PCR products. The remaining four isolates showed no detectable bands and were therefore considered PCR-negative. Overall, the results demonstrate that 60% of the presumptively identified isolates were molecularly confirmed as Helicobacter species using genus-specific PCR.
Molecular confirmations of presumptive H. pylori isolates
PCR amplification using H. pylori-specific primers (HPYF/HPYR) yielded the expected 138 bp product in three of the six Helicobacter genus-positive isolates, confirming them as H. pylori. The molecular size marker showed appropriate reference bands, while the negative control showed no amplification, indicating assay specificity and the absence of contamination. The remaining three isolates showed no detectable 138 bp band and were therefore not confirmed as H. pylori.
Antibiotic susceptibility
All confirmed H. pylori isolates were resistant to ciprofloxacin, azithromycin, and amoxicillin. Complete susceptibility (100%) was observed with streptomycin, which demonstrated strong inhibitory activity against all isolates. Two isolates were susceptible to ofloxacin, while only one isolate showed susceptibility to fluoroquinolones, clarithromycin, and amoxicillin-clavulanate. Gentamicin showed no activity against any of the isolates. Overall, the results indicate the presence of multidrug-resistant H. pylori strains, with streptomycin remaining the most consistently effective agent among those tested.
Discussion
Prevalence of H. pylori infection in relation to age and gender
This study provides important insights into the epidemiology of H. pylori infection among dyspeptic and gastritic patients, highlighting both shared global patterns and distinct local dynamics. The observed prevalence of H. pylori infection was higher among dyspeptic patients (66.7%) than among gastritis patients assessed via stool antigen testing (48%). This difference is clinically meaningful and aligns with the established role of H. pylori as a major etiological agent of dyspeptic symptoms, including chronic gastritis and peptic ulcer disease [26]. The substantial prevalence observed in the broader gastritic population, although lower, further confirms that H. pylori remain a key contributor to gastric pathology, even though other infectious and non-infectious factors may also play a role. These findings support current clinical recommendations advocating a “test-and-treat” strategy in symptomatic patients to enable targeted eradication therapy and reduce unnecessary invasive procedures [27].
Age was significantly associated with H. pylori infection in both patient groups, underscoring the influence of host age on infection dynamics. Among dyspeptic patients, the highest prevalence was recorded in the 30–39-year age group, followed by a progressive decline in older age groups. This pattern likely reflects the natural history of H. pylori infection, whereby acquisition commonly occurs in childhood, but clinical manifestations often become apparent in early to mid-adulthood as cumulative mucosal damage and chronic inflammation progress [28]. The reduced prevalence observed in individuals aged 50–59 years may be partially explained by a birth cohort effect, whereby older individuals experienced lower rates of childhood acquisition due to gradual improvements in hygiene, sanitation, and living conditions over time [29], [30].
In contrast, the gastritic patient cohort exhibited the highest prevalence in the 50–59-year age group, a finding of particular concern given the strong association between long-standing H. pylori infection and severe gastric sequelae, including gastric atrophy, intestinal metaplasia, and gastric carcinoma [31]. The marked decline in prevalence among individuals older than 60 years further supports the birth cohort hypothesis, suggesting reduced exposure in earlier decades. Notably, the secondary peak observed among younger adults (20–29 years) indicates ongoing transmission within the community, highlighting persistent public health challenges related to sanitation, water quality, and overcrowding [32].
Gender-related analysis revealed differing patterns between the two patient groups. The absence of a statistically significant gender difference among dyspeptic patients suggests that susceptibility to symptomatic H. pylori infection may not be inherently sex-dependent. However, the higher prevalence observed among females in the gastritic cohort is noteworthy. While this finding may partly reflect differences in healthcare-seeking behavior, emerging evidence suggests that hormonal factors, particularly estrogen, may influence gastric mucosal immunity and H. pylori colonization density [33]. Additionally, sex-specific variations in gut microbiome composition, which can interact with H. pylori and modulate inflammatory responses, may contribute to the observed disparity [34]. These observations underscore the need for sex-stratified analyses in future epidemiological and mechanistic studies.
Occurrence of H. pylori and other microorganisms in dyspeptic and gastritic patients
Culture-based identification yielded H. pylori in only 10 of the 75 isolates obtained, suggesting a relatively low recovery rate. This finding contrasts with reports of culture positivity rates ranging from 40% to 80% in gastric biopsy samples [35], [36] and with Nigerian studies reporting prevalence rates exceeding 80% using molecular techniques [37], [38]. The lower isolation rate observed in this study may be attributable to several factors, including prior exposure to antibiotics or proton pump inhibitors, delays in sample transport, stringent growth requirements of H. pylori, and inherent limitations of culture-based methods. These challenges reinforce the importance of integrating culture with more sensitive diagnostic tools such as PCR and rapid urease tests to improve detection accuracy [39].
The predominance of E. coli among non-H. pylori isolates is consistent with its status as dominant component of intestinal microbiota and its recognized public health relevance, particularly with respect to extended-spectrum β-lactamase (ESBL)-producing strains [40]. Similarly, the recovery of Pseudomonas species supports growing evidence that Pseudomonas aeruginosa may be more frequently involved in gastrointestinal colonization and disease than previously appreciated [41]. The detection of Bacillus species likely reflects transient environmental contamination, while the presence of yeast cells aligns with recent findings highlighting the gut mycobiome as an integral component of gastrointestinal microbial ecosystems [42].
Molecular identification of Helicobacter species and H. pylori
Molecular analysis revealed notable discrepancies between phenotypic identification and PCR confirmation, with only 6 of 10 presumptive Helicobacter isolates confirmed at the genus level. This finding underscores the limited specificity of culture and biochemical methods and highlights the superior accuracy of PCR-based diagnostics for Helicobacter detection [43]. Furthermore, only half of the PCR-confirmed Helicobacter isolates were identified as H. pylori, reflecting the diversity of the Helicobacter genus and the potential involvement of non-pylori species such as H. heilmannii and H. felis in gastric colonization and disease [44]. These findings emphasize the need for precise molecular characterization to fully elucidate the epidemiology and pathogenic relevance of Helicobacter species in clinical settings.
Antibiotic susceptibility patterns
The antibiotic susceptibility profiles reveal concerning resistance trends across multiple antibiotic classes. Resistance to fluoroquinolones, including ciprofloxacin and ofloxacin, was prominent, consistent with global reports linking widespread antibiotic use to increasing fluoroquinolone resistance in H. pylori [45]. Similarly, high resistance rates to macrolides such as clarithromycin and azithromycin are particularly troubling, given the central role of these agents in standard eradication regimens. These findings corroborate reports of declining clarithromycin efficacy and underscore the need for resistance-guided therapy [46], [47].
Beta-lactam antibiotics demonstrated variable susceptibility patterns, suggesting heterogeneity in resistance mechanisms among circulating strains. Such variability highlights the importance of local antimicrobial surveillance to inform empirical treatment decisions and optimize eradication outcomes. In contrast, aminoglycosides, particularly streptomycin, showed high efficacy against the tested isolates, consistent with previous studies reporting low resistance rates to this class [48]. Sulfonamides exhibited moderate activity, indicating potential utility in selected cases, although susceptibility testing remains essential to ensure therapeutic success. Overall, the wide variation in zones of inhibition observed across isolates reflects substantial strain-specific differences in resistance profiles. This heterogeneity underscores the growing challenge posed by multidrug-resistant H. pylori and reinforces the need for routine susceptibility testing and tailored treatment strategies to improve eradication rates and limit further resistance development.
Public health interventions aimed at improving sanitation, access to clean water, and awareness of H. pylori transmission routes are also recommended to reduce infection rates.
Study limitations
This study has several limitations that should be considered when interpreting the findings. First, the relatively small number of H. pylori isolates confirmed by molecular methods and subjected to antibiotic susceptibility testing may limit the generalizability of the resistance patterns observed. Second, reliance on culture-based identification may have underestimated the true prevalence of H. pylori, given its fastidious growth requirements and the potential effects of prior antibiotic or proton pump inhibitor use among participants. Third, the cross-sectional design of the study precludes assessment of causal relationships, temporal trends, and treatment outcomes. In addition, virulence determinants and specific resistance-associated gene mutations were not investigated, which limits insights into strain pathogenicity and molecular mechanisms underlying antimicrobial resistance.
Based on the findings, larger multicenter studies incorporating increased sample sizes are recommended to provide more representative prevalence and resistance data. The integration of advanced molecular techniques, including detection of virulence genes and resistance-associated mutations, would enhance understanding of strain diversity and therapeutic failure. Finally, longitudinal studies evaluating treatment outcomes and reinfection rates are needed to inform optimized management strategies and policy development.
Notes
Author’s ORCID
Ngene AC: https://orcid.org/0000-0003-4730-2834
Ethical approval
Ethical clearance was obtained from the Ethics Committee of the Jos University Teaching Hospital.
Funding
This research was supported by the University of Jos through the Institutional-Based Research (IBR) Grant, Grant No. 036.
Competing interests
The authors declare that they have no competing interests.
References
[1] Egbere OJ, Emmanuel AU, Okopi JAA, Ali MA, Okojokwu OJ, Fowora M, Danladi MM, Ngene AC. Inhibitory effects of Lactobacillus acidophilus and Lactobacillus casei isolated from "Kunun zaki" (a Nigerian fermented beverage) against Helicobacter pylori. World J Microbiol. 2020;5(1):160-7.[2] Elbehiry A, Marzouk E, Aldubaib M, Abalkhail A, Anagreyyah S, Anajirih N, Almuzaini AM, Rawway M, Alfadhel A, Draz A, Abu-Okail A. Helicobacter pylori Infection: Current Status and Future Prospects on Diagnostic, Therapeutic and Control Challenges. Antibiotics (Basel). 2023 Jan 17;12(2):191. DOI: 10.3390/antibiotics12020191
[3] Duan M, Li Y, Liu J, Zhang W, Dong Y, Han Z, Wan M, Lin M, Lin B, Kong Q, Ding Y, Yang X, Zuo X, Li Y. Transmission routes and patterns of helicobacter pylori. Helicobacter. 2023 Feb;28(1):e12945. DOI: 10.1111/hel.12945
[4] Shafiq H, Zafar I, Shafiq S. A systematic review on current trends of socioeconomic disparities and Helicobacter pylori associated risk factors. Res Med Sci Rev. 2024;2(3):1732-48.
[5] Joy OS, Aluko EF, Ayanbolade OR, Olowe OA. Public health perspectives of zoonotic potential of H. pylori. Adv Anal Pathol. 2025 Mar;1:22-38. DOI: 10.17582/journal.ppa/2025/1.22.38
[6] Ansari S, Yamaoka Y. Survival of Helicobacter pylori in gastric acidic territory. Helicobacter. 2017 Aug;22(4):10.1111/hel.12386. DOI: 10.1111/hel.12386
[7] Cheok YY, Lee CYQ, Cheong HC, Vadivelu J, Looi CY, Abdullah S, Wong WF. An Overview of Helicobacter pylori Survival Tactics in the Hostile Human Stomach Environment. Microorganisms. 2021 Dec 3;9(12):2502. DOI: 10.3390/microorganisms9122502
[8] Smith SI, Ajayi A, Jolaiya T, Onyekwere C, Setshedi M, Schulz C, Otegbayo JA, Ndip R, Dieye Y, Alboraie M, Ally R, Gunturu R, Hyasinta J, Ugiagbe R, Ndububa D, Arigbabu A; African Helicobacter and Microbiota Study Group. Helicobacter pylori Infection in Africa: Update of the Current Situation and Challenges. Dig Dis. 2022;40(4):535-44. DOI: 10.1159/000518959
[9] Emmanuel BN, Peter DA, Peter MO, Adedayo IS, Olaifa K. Helicobacter pylori infection in Africa: comprehensive insight into its pathogenesis, management, and future perspectives. J Umm Al-Qura Univ Appl Sci. 2025 Jun;11(2):378-401. DOI: 10.1007/s43994-024-00166-6
[10] Hatta MNA, Mohamad Hanif EA, Chin SF, Neoh HM. Pathogens and Carcinogenesis: A Review. Biology (Basel). 2021 Jun;10(6):533. DOI: 10.3390/biology10060533
[11] Bakhti SZ, Latifi-Navid S, Safaralizadeh R. Helicobacter pylori-related risk predictors of gastric cancer: The latest models, challenges, and future prospects. Cancer Med. 2020 Jul;9(13):4808-22. DOI: 10.1002/cam4.3068
[12] Ali A, AlHussaini KI. Helicobacter pylori: A Contemporary Perspective on Pathogenesis, Diagnosis and Treatment Strategies. Microorganisms. 2024 Jan 22;12(1):222. DOI: 10.3390/microorganisms12010222
[13] Ansari S, Yamaoka Y. Helicobacter pylori Infection, Its Laboratory Diagnosis, and Antimicrobial Resistance: a Perspective of Clinical Relevance. Clin Microbiol Rev. 2022 Sep;35(3):e0025821. DOI: 10.1128/cmr.00258-21
[14] Hu Y, Zhu Y, Lu NH. Novel and Effective Therapeutic Regimens for Helicobacter pylori in an Era of Increasing Antibiotic Resistance. Front Cell Infect Microbiol. 2017 May 5;7:168. DOI: 10.3389/fcimb.2017.00168
[15] Kuve SA. Screening for Carriage and Antimicrobial Resistance of Helicobacter pylori, Escherichia coli and Klebsiella species among Patients presenting with Gastritis in Kibera and Dagoretti areas in Nairobi, Kenya [Dissertation]. 2023.
[16] Dossouvi KM, Sellera FP, Ibadin EE, Gnagnon KW, Elkelish A. Recommendations and Adjuvant Therapies to Mitigate Clarithromycin-Resistant Helicobacter pylori-Associated Infections in Africa. Infect Drug Resist. 2025 Jun 26;18:3169-74. DOI: 10.2147/IDR.S528422
[17] Smith SI, Fowora MA, Otegbayo JA, Abdulkareem FB, Omonigbehin EA, Adegboyega A, Contreras M, Haas R. Comparison of PCR with other diagnostic techniques for the detection of H. pylori infection in patients presenting with gastroduodenal symptons in Nigeria. Int J Mol Epidemiol Genet. 2011;2(2):178-84.
[18] Fatima A, Ibrahim M, Naseer A, Pervez A, Asad M, Shah AA, Hasan F, Alonazi WB, Ferheen I, Khan S. Identification, Genome Sequencing, and Characterizations of Helicobacter pylori Sourced from Pakistan. Microorganisms. 2023 Oct 29;11(11):2658. DOI: 10.3390/microorganisms11112658
[19] Coudron PE, Kirby DF. Comparison of rapid urease tests, staining techniques, and growth on different solid media for detection of Campylobacter pylori. J Clin Microbiol. 1989 Jul;27(7):1527-30. DOI: 10.1128/jcm.27.7.1527-1530.1989
[20] Lee N, Lee TT, Fang KM. Assessment of four rapid urease test systems for detection of Helicobacter pylori in gastric biopsy specimens. Diagn Microbiol Infect Dis. 1994 Feb;18(2):69-74. DOI: 10.1016/0732-8893(94)90067-1
[21] Kelly SM, Pitcher MC, Farmery SM, Gibson GR. Isolation of Helicobacter pylori from feces of patients with dyspepsia in the United Kingdom. Gastroenterology. 1994 Dec;107(6):1671-4. DOI: 10.1016/0016-5085(94)90806-0
[22] Mujtaba A, Masud T, Ahmad A, Naqvi SMS, Ismail A, Tahir MI, Yousaf AA, Ibrahim MS. Isolation and antimicrobial susceptibility testing of Helicobacter pylori strains from gastric biopsies from Pakistani patients. Pak J Pharm Sci. 2019 Sep;32(5(Supplementary)):2279-85.
[23] Cheesbrough M. District laboratory practice in tropical countries, part 2. Cambridge university press;2005. DOI: 10.1017/CBO9780511581304
[24] Ngene AC, Ohaegbu CG, Awom IE, Egbere JO, Onyimba IA, Coulthard OD, Umar U, Ohaeri UC, Nnadi NN, Aguiyi JC. High prevalence of multidrug resistant enterobacteriaceae isolated from wastewater and soil in Jos Metropolis, Plateau State, Nigeria. African J Bacteriol Res. 2021 Dec 31;13(2):22-9. DOI: 10.5897/JBR2021.0336
[25] Hemen WN, Ohaegbu CG, Onyimba IA, Odu CE, Ngene AC, Danladi MM, Dashen MM, Ochiheche EG, Entonu EE, Ogaji AO, Egbere OJ. Antibiotic susceptibility profile of bacterial pathogens and health hazards associated with municipal solid wastes in Gboko Town, Nigeria. J Adv Microbiol. 2022:132-46. DOI: 10.9734/JAMB/2022/v22i10676
[26] Malfertheiner P, Megraud F, Rokkas T, Gisbert JP, Liou JM, Schulz C, Gasbarrini A, Hunt RH, Leja M, O'Morain C, Rugge M, Suerbaum S, Tilg H, Sugano K, El-Omar EM; European Helicobacter and Microbiota Study group. Management of Helicobacter pylori infection: the Maastricht VI/Florence consensus report. Gut. 2022 Aug 8:gutjnl-2022-327745. DOI: 10.1136/gutjnl-2022-327745
[27] Namikawa K, Purisevic FL, Thorsteinsson JB, Bjornsson ES. Helicobacter pylori Across Continents: Contrasts in Epidemiology, Genetics, Clinical Impact, and Management Between East and West. Int J Mol Sci. 2025 Nov 25;26(23):11408. DOI: 10.3390/ijms262311408
[28] Ren S, Cai P, Liu Y, Wang T, Zhang Y, Li Q, Gu Y, Wei L, Yan C, Jin G. Prevalence of Helicobacter pylori infection in China: A systematic review and meta-analysis. J Gastroenterol Hepatol. 2022 Mar;37(3):464-70. DOI: 10.1111/jgh.15751
[29] Leung WK, Wong IOL, Cheung KS, Yeung KF, Chan EW, Wong AYS, Chen L, Wong ICK, Graham DY. Effects of Helicobacter pylori Treatment on Incidence of Gastric Cancer in Older Individuals. Gastroenterology. 2018 Jul;155(1):67-75. DOI: 10.1053/j.gastro.2018.03.028
[30] Li Y, Choi H, Leung K, Jiang F, Graham DY, Leung WK. Global prevalence of Helicobacter pylori infection between 1980 and 2022: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol. 2023 Jun;8(6):553-64. DOI: 10.1016/S2468-1253(23)00070-5
[31] Sugano K, Tack J, Kuipers EJ, Graham DY, El-Omar EM, Miura S, Haruma K, Asaka M, Uemura N, Malfertheiner P; faculty members of Kyoto Global Consensus Conference. Kyoto global consensus report on Helicobacter pylori gastritis. Gut. 2015 Sep;64(9):1353-67. DOI: 10.1136/gutjnl-2015-309252
[32] Zamani M, Ebrahimtabar F, Zamani V, Miller WH, Alizadeh-Navaei R, Shokri-Shirvani J, Derakhshan MH. Systematic review with meta-analysis: the worldwide prevalence of Helicobacter pylori infection. Aliment Pharmacol Ther. 2018 Apr;47(7):868-76. DOI: 10.1111/apt.14561
[33] Wang J, Xu L, Shi R, Huang X, Li SW, Huang Z, Zhang G. Gastric atrophy and intestinal metaplasia before and after Helicobacter pylori eradication: a meta-analysis. Digestion. 2011;83(4):253-60. DOI: 10.1159/000280318
[34] Peng C, Li X, Li Y, Xu X, Ouyang Y, Li N, Lu N, Zhu Y, He C. Sex-specific effects of gastrointestinal microbiome disruptions on Helicobacter pylori-induced gastric carcinogenesis in INS-GAS mice. Biol Sex Differ. 2025 Feb;16(1):15. DOI: 10.1186/s13293-025-00700-z
[35] Sousa C, Ferreira R, Santos SB, Azevedo NF, Melo LDR. Advances on diagnosis of Helicobacter pylori infections. Crit Rev Microbiol. 2023 Nov;49(6):671-92. DOI: 10.1080/1040841X.2022.2125287
[36] Ramis IB, de Moraes EP, Fernandes MS, Mendoza-Sassi R, Rodrigues O, Juliano CR, Scaini CJ, da Silva PE. Evaluation of diagnostic methods for the detection of Helicobacter pylori in gastric biopsy specimens of dyspeptic patients. Braz J Microbiol. 2012 Jul;43(3):903-8. DOI: 10.1590/S1517-83822012000300008
[37] Ajayi A, Jolaiya T, Smith SI. Direct detection of Helicobacter pylori from biopsies of patients in Lagos, Nigeria using real-time PCR-a pilot study. BMC Res Notes. 2021 Mar;14(1):90. DOI: 10.1186/s13104-021-05505-y
[38] Smith SI, Ajayi A, Jolaiya TF, Essiet U. Prevalence, diagnosis and treatment of Helicobacter pylori infection in Nigeria. Nigerian Journal of Gastroenterology and Hepatology. 2022 Jan 1;14(1):2-10. DOI: 10.4103/njgh.njgh_10_22
[39] Costa LCMC, das Graças Carvalho M, La Guárdia Custódio Pereira AC, Teixeira Neto RG, Andrade Figueiredo LC, Barros-Pinheiro M. Diagnostic Methods for Helicobacter pylori. Med Princ Pract. 2024;33(3):173-84. DOI: 10.1159/000538349
[40] Gudrā D. The long-term impact of Helicobacter pylori eradication therapy on the human gastrointestinal microbiome and extended-spectrum β-lactamase (ESBL) coding gene levels [Dissertation]. Latvijas University;2024.
[41] Pettigrew MM, Gent JF, Kong Y, Halpin AL, Pineles L, Harris AD, Johnson JK. Gastrointestinal Microbiota Disruption and Risk of Colonization With Carbapenem-resistant Pseudomonas aeruginosa in Intensive Care Unit Patients. Clin Infect Dis. 2019 Aug;69(4):604-13. DOI: 10.1093/cid/ciy936
[42] Richard ML, Sokol H. The gut mycobiota: insights into analysis, environmental interactions and role in gastrointestinal diseases. Nat Rev Gastroenterol Hepatol. 2019 Jun;16(6):331-45. DOI: 10.1038/s41575-019-0121-2
[43] Cardos AI, Maghiar A, Zaha DC, Pop O, Fritea L, Miere Groza F, Cavalu S. Evolution of Diagnostic Methods for Helicobacter pylori Infections: From Traditional Tests to High Technology, Advanced Sensitivity and Discrimination Tools. Diagnostics (Basel). 2022 Feb 16;12(2):508. DOI: 10.3390/diagnostics12020508
[44] Taillieu E, De Witte C, De Schepper H, Van Moerkercke W, Rutten S, Michiels S, Arnst Y, De Bruyckere S, Francque S, van Aert F, George C, Callewaert E, Callewaert T, Vanneste G, Vanderstraeten E, Van Heddegem N, Vansteelant M, Chiers K, Haesebrouck F, Van Steenkiste C. Clinical significance and impact of gastric non-Helicobacter pylori Helicobacter species in gastric disease. Aliment Pharmacol Ther. 2023 Jun;57(12):1432-44. DOI: 10.1111/apt.17488
[45] Yu Y, Xue J, Lin F, Liu D, Zhang W, Ru S, Jiang F. Global Primary Antibiotic Resistance Rate of Helicobacter pylori in Recent 10?years: A Systematic Review and Meta-Analysis. Helicobacter. 2024;29(3):e13103. DOI: 10.1111/hel.13103
[46] Gisbert JP. Empirical or susceptibility-guided treatment for Helicobacter pylori infection? A comprehensive review. Therap Adv Gastroenterol. 2020 Nov 12;13:1756284820968736. DOI: 10.1177/1756284820968736
[47] Muttiah B, Wahid W, Sukri A, Hanafiah A. Towards Effective Helicobacter pylori Eradication: Emerging Therapies in the Wake of Antibiotic Resistance. Int J Mol Sci. 2025 Jun 24;26(13):6064. DOI: 10.3390/ijms26136064
[48] Ng HY, Leung WK, Cheung KS. Antibiotic Resistance, Susceptibility Testing and Stewardship in Helicobacter pylori Infection. Int J Mol Sci. 2023 Jul 20;24(14):11708. DOI: 10.3390/ijms241411708




